| Original Literature | Model OverView |
|---|---|
|
Publication
Title
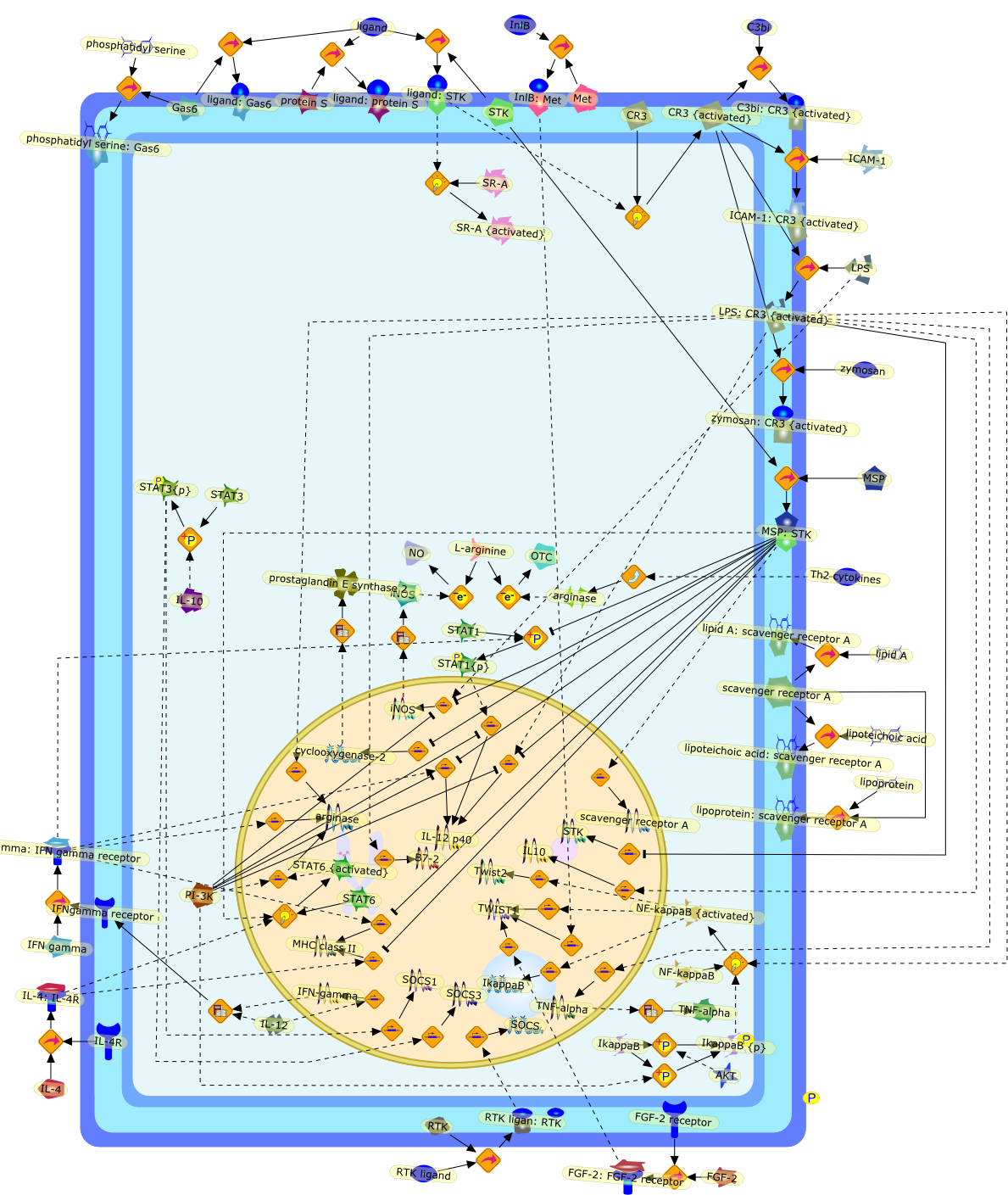
Receptor tyrosine kinases and the regulation of macrophage activation.
Affiliation
Department of Veterinary Science, The Pennsylvania State University, UniversityPark, PA 16802-3500, USA. phc7@psu.edu
Abstract
PMID
14726496
|