| Original Literature | Model OverView |
|---|---|
|
Publication
Title
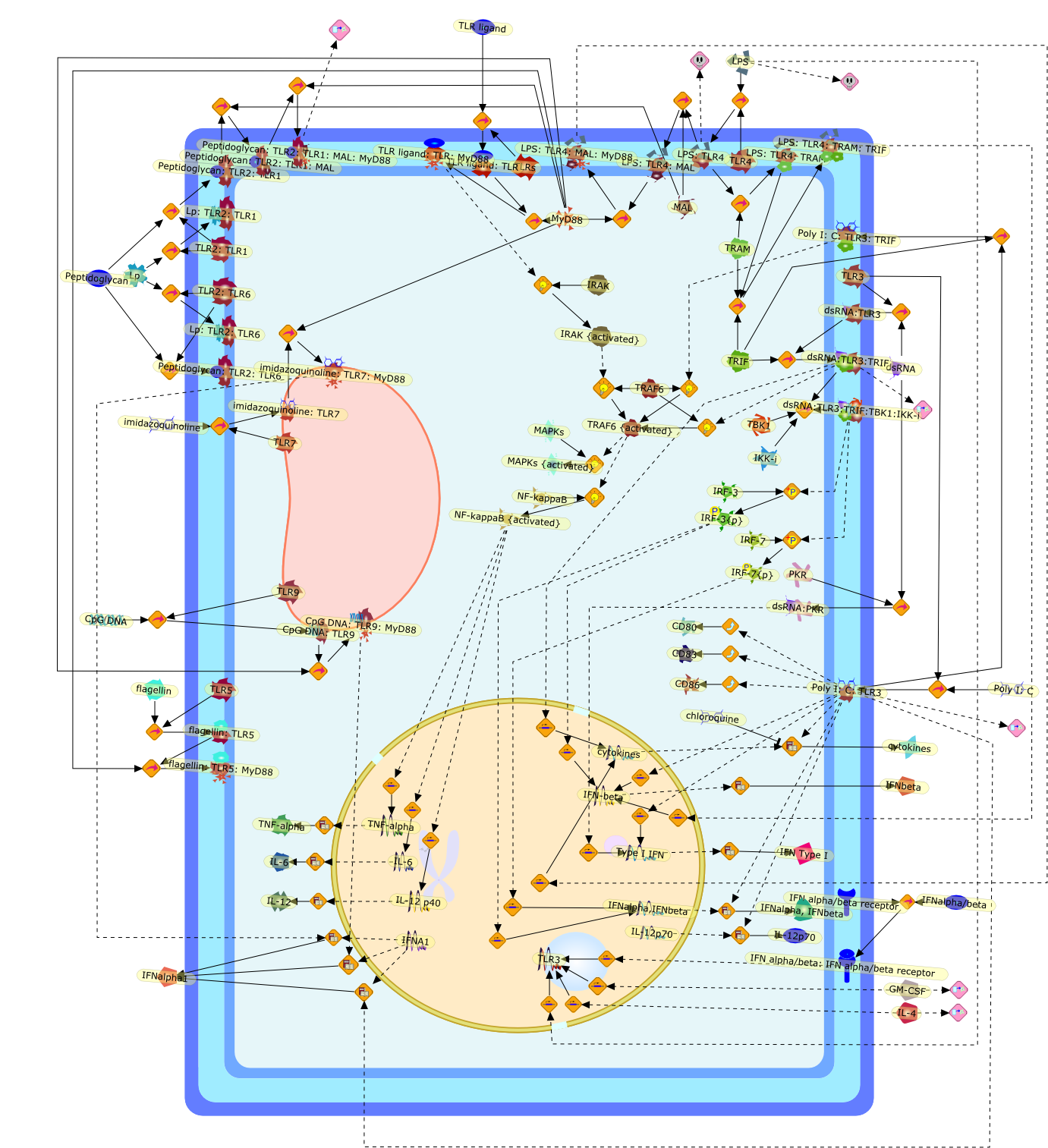
Toll-like receptor 3: a link between toll-like receptor, interferon and viruses.
Affiliation
Department of Immunology, Osaka Medical Center for Cancer and CardiovascularDiseases, Osaka, Japan. matsumoto-mi@mc.pref.osaka.jp
Abstract
Production of type I interferon (IFN-alpha/beta) by virus-infected cells is thecentral event in their antiviral immune responses. In mammalian cells,IFN-alpha/beta gene transcription is induced through distinct signaling pathwaysby viral infection or by treatment with double-stranded (ds) RNA, which is anintermediate of virus replication. Toll-like receptor 3 (TLR3) was found torecognize dsRNA and transmit signals to activate NF-kappaB and the IFN-betapromoter. Recent identification of the TLR3-adaptor protein and its downstreamsignaling molecules, which are involved in IFN-alpha/beta production, revealed anovel IFN-inducing pathway for an anti-viral immune response. Here, we summarizethe current knowledge of TLR3-mediated immune responses.
PMID
15031527
|